Calculates the average absorbance and fluorescence values given multiple check standards to create a long-term standard that the daily check standard can be checked against for consistency.
Usage
create_std(
dir,
method = "default",
meta_name = NULL,
sheet = NULL,
abs_pattern = "Abs",
iblank = "BEM",
type = "eem",
recursive = FALSE,
qaqc_dir = get_qaqc_dir(),
update_config = TRUE
)Source
Hansen, A. M., Fleck, J., Kraus, T. E. C., Downing, B. D., von Dessonneck, T., & Bergamaschi, B. (2018). Procedures for using the Horiba Scientific Aqualog® fluorometer to measure absorbance and fluorescence from dissolved organic matter (USGS Numbered Series No. 2018-1096). U.S. Geological Survey. doi:10.3133/ofr20181096
Arguments
- dir
Path to a folder containing long-term EEMs and/or absorbance files. All files in this directory will be loaded.
- method
A character string describing the method associated with the MDL files.
- meta_name
Name of the metadata file. Optional if the metadata file is the only
.xlsxor.csvfile indir. If not specified, the function attempts to find a single metadata file and errors if multiple files are present.- sheet
Name of the sheet containing metadata (only required if the metadata is not on the first sheet of an
.xlsxfile).- abs_pattern
A character string containing a base::regular expression used to identify absorbance files.
- iblank
Optional. A character string containing a base::regular expression used to identify instrument blanks.
- type
Which MDL to calculate: either "eem" or "abs".
- recursive
Logical. Should the function recursively search directories?
- qaqc_dir
Directory in which to save the QAQC
.rdsfile. Default: The QAQC directory in the current package environment, usually the user configured QAQC directory. IfNA, the function returns the MDL object instead of saving it.- update_config
Logical. Should function ask to update user_config file with the default QAQC directory location?
Value
If
dir = FALSE: aneemorabsobject containing the averaged check standard values.Otherwise: saves an
.rdsfile containing the averaged check standard and invisibly returns the file path.
Details
To ensure optical measurements are consistent across days, Hansen et al. (2018) recommend using a standard reference material: Pure Leaf® unsweetened black tea. The tea standard should be diluted to 1% concentration before analysis. It is recommended to run the tea standard at the start and end of each analysis run. According to USGS standards, tea standard indices measured on a given day should fall within 20% of the long-term average (Hansen et al. 2018).
To calculate the average check standard you need:
A directory containing check standards with their associated instrument blanks (less than 20 will prompt a warning)
Note: sample names must be unique
Metadata for the check samples including (at a minimum) the integration time and raman area in a single file, formatted as a metadata file (see metadata)
Examples
eem_std <- create_std(file.path(system.file("extdata", package = "eemanalyzeR"),"long-term-std"),
meta_name="longterm-checkstd-metadata.csv", abs_pattern = "ABS",
type="eem", qaqc_dir = NA, update_config=FALSE)
#> NOTE: removed previous 'readme' file
#> Warning: Calculating average check standard based on less than 20 samples, average may be unreliable
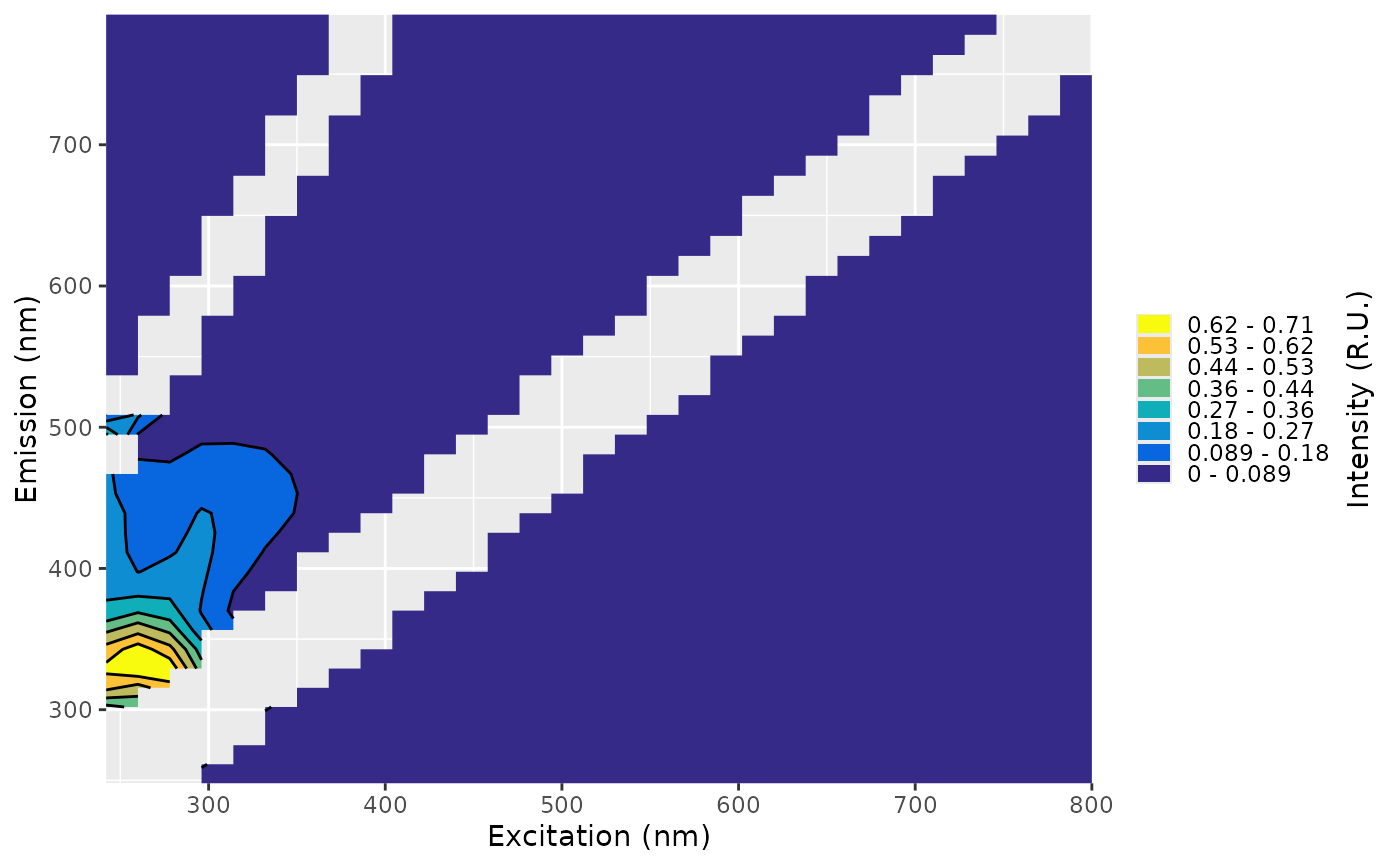
plot(eem_std)